NodeFills <- mapVisualProperty('node fill color','node_attributes','d',c("A","B"), c("#FF9900","#66AAAA"))ĪrrowShapes <- mapVisualProperty('Edge Target Arrow Shape','interaction','d',c("activates","inhibits","interacts"),c("Arrow","T","None"))ĮdgeWidth <- mapVisualProperty('edge width','weight','p')ĬreateVisualStyle(style. NodeLabels <- mapVisualProperty('node label','id','p') SetVisualStyle('Marquee') set up my own style Node <- lim("merge_CytoscapeInput-nodes-paleturquoise.txt")Ĭolnames(node) it need to modify colnamesĬolnames(node) <- c("id", "altName", "node_attributes")ĬreateNetworkFromDataFrames(node,edge, title="paleturquoise network", collection="DataFrame paleturquoise") You can resize the columns by hovering between the column headers and clicking and dragging on the resize cursor that appears. # use the connection with Cytoscape through the 'RC圓' packageĮdge <- lim("merge_CytoscapeInput-edges-paleturquoise.txt")Ĭolnames(edge) it need to modify colnamesĬolnames(edge) <- c("source", "target", "weight", "direction","fromAltName", "toAltName") Table styles are now part of the Network Style and are applied to the node and edge tables. 
Edge label rotation allows edge labels to be aligned with the edge slope. NodeName altName nodeAttrĬytoscapePing () # make sure cytoscape is openĬytoscapeVersionInfo () cytoscapeVersion "3.10.0" Hold Ctrl-Shift on Windows/Linux, or Cmd-Shift on Mac, then use the mouse to draw a free-form loop around nodes you wish to select. Make use of Cytoscape 3.3.2 (edge point fix, cytoscape/cytoscape.js2250) 0.3.0- Jan 14, 2019. With Dash, you can build biological networks, social networks, informational networks, and pretty much. Platform: x86_64-w64-mingw32/圆4 (64-bit)įromNode toNode weight direction fromAltName toAltName 2,348 views We'll dive deeper into the Styling chapter of Dash Cytoscape. Cytoscape is a free software package for visualizing, modeling and analyzing molecular and genetic interaction networks. I get the same degree for the whole table and I don't understand if I'm doing. Then I do the network analysis to get the 'degree', but something is strange. The core in the styles is for the viewport/background as. Hello, I got the network with WGCN, use RC圓 to import the networks to cytoscape. I guess that selection-box-color, selection-box-border-color and selection-box-border-width allow to change the styles. while pressing the mouse over the node/edge, not after (see attached images). R version 4.2.3 ( ucrt) - "Shortstop Beagle"Ĭopyright (C) 2023 The R Foundation for Statistical Computing Im wondering how to change the node and edges styles when selecting them, i.e. 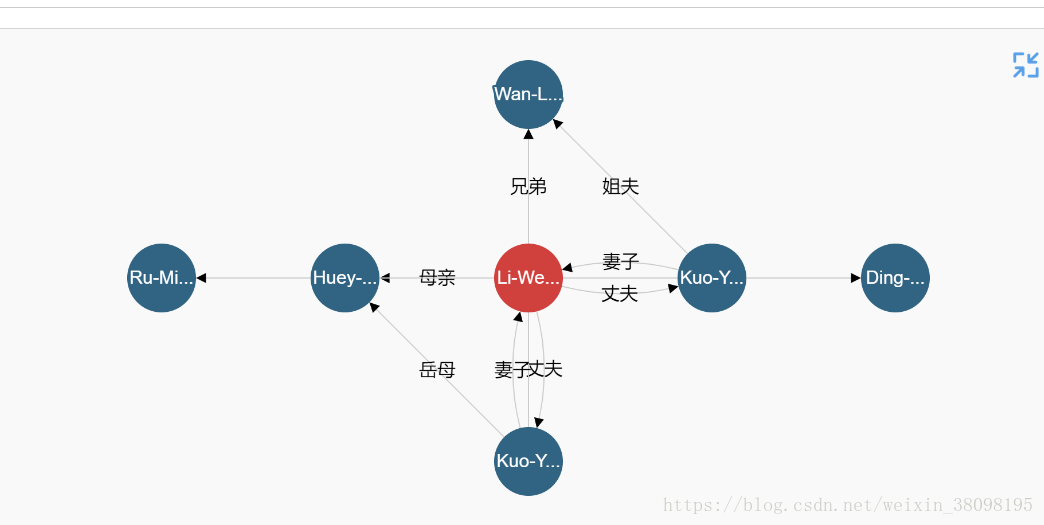
I get the same degree for the whole table and I don't understand if I'm doing something wrong in cytoscape or in the R studio code. Then I do the network analysis to get the "degree", but something is strange. I got the network with WGCN, use RC圓 to import the networks to cytoscape.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
